Loop and domain calling#
This notebook demonstrates how to identify chromatin loops and TADs from the seqFISH+ dataset from Takei et al, 2021. The 25Kb subset used here can be downloaded from the 4DN data portal with file IDs: 4DNFIHF3JCBY (biological replicate 1) and 4DNFIQXONUUH (biological replicate 2).
[1]:
import matplotlib.pyplot as plt
import arcfish as af
Loading FOF_CT-core file#
arcfish.pp.FOF_CT_Loader is the base class to read in FOF_CT files. The input can be a string, a list, or a dictionary like here. It can also read in other FOF_CT files other than FOF_CT-core (check api documentation here).
For FOF_CT-core files, loader.chr_ids show a list of available chromosomes.
[2]:
loader = af.pp.FOF_CT_Loader({
"rep1": "4DNFIHF3JCBY.csv", "rep2": "4DNFIQXONUUH.csv"
}, nm_ratio={c: 1000 for c in "XYZ"})
loader.chr_ids
[2]:
array(['chr1', 'chr2', 'chr3', 'chr4', 'chr5', 'chr6', 'chr7', 'chr8',
'chr9', 'chr10', 'chr11', 'chr12', 'chr13', 'chr14', 'chr15',
'chr16', 'chr17', 'chr18', 'chr19', 'chrX'], dtype=object)
The create_adata function creates an AnnData object for a specific chromosome with the following fields:
n_obs: the number of chromatin tracesn_vars: the number of imaging lociX,Y, andZ: 3D coordinatesChrom_StartandChrom_End: 1D genomic location of each locusChrom: chromosome number (“chr6” in this case)
[3]:
adata = loader.create_adata("chr6")
adata
[3]:
AnnData object with n_obs × n_vars = 921 × 60
var: 'Chrom_Start', 'Chrom_End'
uns: 'Chrom'
layers: 'X', 'Y', 'Z'
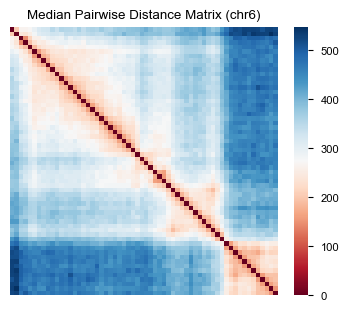
We can calculate the median pairwise distance matrix and visualize it using af.pp.median_pdist and af.pl.pairwise_heatmap.
[4]:
af.pp.median_pdist(adata, inplace=True)
adata
[4]:
AnnData object with n_obs × n_vars = 921 × 60
var: 'Chrom_Start', 'Chrom_End'
uns: 'Chrom'
layers: 'X', 'Y', 'Z'
varp: 'med_dist'
[5]:
fig, ax = plt.subplots(figsize=(4, 3))
af.pl.pairwise_heatmap(adata.varp["med_dist"], ax=ax, cmap="RdBu",
title="Median Pairwise Distance Matrix (chr6)")
Loop calling#
First we filter and normalize the data. This appends ‘med_dist’, ‘raw_var_X’, ‘raw_var_Y’, ‘raw_var_Z’, ‘count_X’, ‘count_Y’, ‘count_Z’, ‘var_X’, ‘var_Y’, ‘var_Z’ to the anndata object, where var_* are the axis variance matrices.
[6]:
af.pp.filter_normalize(adata)
adata
[6]:
AnnData object with n_obs × n_vars = 921 × 60
var: 'Chrom_Start', 'Chrom_End'
uns: 'Chrom'
layers: 'X', 'Y', 'Z'
varp: 'med_dist', 'raw_var_X', 'raw_var_Y', 'raw_var_Z', 'count_X', 'count_Y', 'count_Z', 'var_X', 'var_Y', 'var_Z'
Create a LoopCaller object with an FDR cutoff of 0.1. The call_loops() method returns a dictionary containing the analysis results.
[7]:
caller = af.tl.LoopCaller(fdr_cutoff=0.1)
loop_result = caller.call_loops(adata)
loop_result.keys()
[7]:
dict_keys(['axis_stat', 'axis_pval', 'stat', 'pval', 'fdr', 'candidate', 'label', 'summit', 'final'])
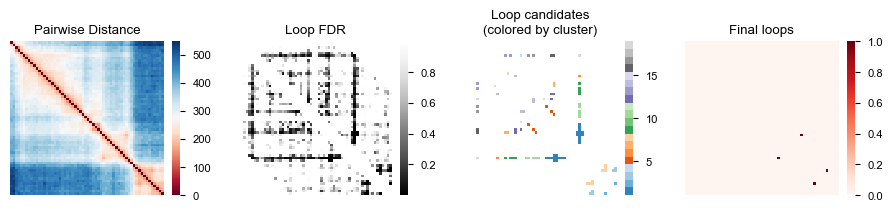
Now we create four heatmaps to visualize the loop calling process: the original distance matrix, FDR values, loop candidates grouped by clusters, and the final loop calls.
[8]:
fig, axes = plt.subplots(1, 4, figsize=(9, 2))
af.pl.pairwise_heatmap(adata.varp["med_dist"], ax=axes[0], cmap="RdBu",
title="Pairwise Distance")
af.pl.pairwise_heatmap(loop_result["fdr"], ax=axes[1], cmap="Greys_r",
title="Loop FDR")
af.pl.pairwise_heatmap(loop_result["label"], ax=axes[2], cmap="tab20c",
title="Loop candidates\n(colored by cluster)")
af.pl.pairwise_heatmap(loop_result["final"], ax=axes[3], cmap="Reds",
title="Final loops")
Convert the loop results to BEDPE format using the to_bedpe() method, then filter for final loops and display the first few rows.
[9]:
bedpe = caller.to_bedpe(loop_result, adata)
bedpe[bedpe["final"]].head()
[9]:
| c1 | s1 | e1 | c2 | s2 | e2 | stat | pval | fdr | candidate | label | summit | final | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1502 | chr6 | 50300000 | 50325000 | chr6 | 50525000 | 50550000 | 7.145024e+09 | 4.454981e-11 | 6.058774e-08 | True | 1.0 | True | True |
| 1729 | chr6 | 50675000 | 50700000 | chr6 | 50800000 | 50825000 | 1.741511e+06 | 1.827780e-07 | 6.214450e-05 | True | 8.0 | True | True |
TAD calling#
Create a TADCaller object with FDR cutoff 0.1 and run call_tads() on the data. This returns a DataFrame with the TAD calling results.
Note here the data is already filtered and normalized. If the data is not preprocessed yet, run filter_normalize first.
[10]:
caller = af.tl.TADCaller(fdr_cutoff=0.1)
tad_result = caller.call_tads(adata)
tad_result.head()
[10]:
| c1 | Chrom_Start | Chrom_End | stat_x | stat_y | stat_z | pval_x | pval_y | pval_z | stat | stat_prox | pval | fdr | fdr_peak | raw_peak | peak | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | chr6 | 49375000 | 49400000 | NaN | NaN | NaN | NaN | NaN | NaN | NaN | NaN | NaN | NaN | False | False | False |
| 1 | chr6 | 49400000 | 49425000 | NaN | NaN | NaN | NaN | NaN | NaN | NaN | NaN | NaN | NaN | False | False | False |
| 2 | chr6 | 49450000 | 49475000 | 0.976542 | 0.986278 | 0.932193 | 0.253538 | 3.494223e-01 | 2.560919e-02 | 4.802816e+00 | 0.018384 | 6.534212e-02 | 1.829579e-01 | False | False | False |
| 3 | chr6 | 49475000 | 49500000 | 0.959849 | 0.917475 | 0.909965 | 0.086188 | 2.053673e-03 | 9.127656e-04 | 1.714311e+02 | 0.036911 | 1.856759e-03 | 8.664876e-03 | True | False | False |
| 4 | chr6 | 49500000 | 49525000 | 0.889484 | 0.811094 | 0.861914 | 0.000053 | 2.063632e-12 | 4.926662e-07 | 4.873805e+10 | 0.078701 | 6.530998e-12 | 6.095598e-11 | True | True | True |
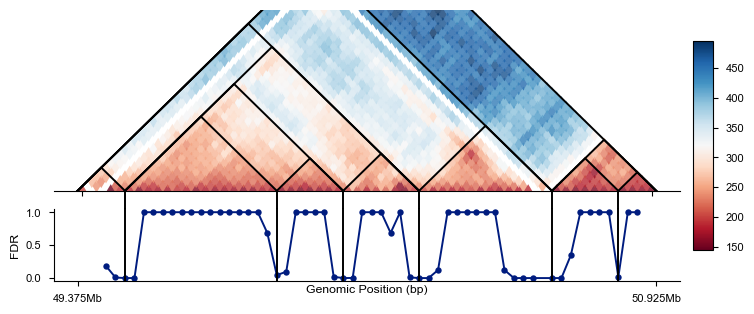
Visualize the TAD boundaries using triangle_domain_boundary(), which creates a triangle plot showing the domain structure.
[11]:
fig = plt.figure(figsize=(7, 2.8))
ax, cbar, cax = af.pl.triangle_domain_boundary(adata, caller, fig=fig)
Convert the TAD results to BEDPE format and display the first few rows.
[12]:
caller.to_bedpe(tad_result).head()
[12]:
| c1 | s1 | e1 | c2 | s2 | e2 | stat1 | stat2 | level | idx1 | idx2 | |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | chr6 | 49375000 | 49400000 | chr6 | 49500000 | 49525000 | NaN | 4.873805e+10 | 0 | 0 | 4 |
| 1 | chr6 | 49500000 | 49525000 | chr6 | 49900000 | 49925000 | 4.873805e+10 | 2.508111e+01 | 0 | 4 | 20 |
| 2 | chr6 | 49900000 | 49925000 | chr6 | 50075000 | 50100000 | 2.508111e+01 | 1.021685e+08 | 0 | 20 | 27 |
| 3 | chr6 | 50075000 | 50100000 | chr6 | 50275000 | 50300000 | 1.021685e+08 | 5.160219e+15 | 0 | 27 | 35 |
| 4 | chr6 | 50275000 | 50300000 | chr6 | 50625000 | 50650000 | 5.160219e+15 | 1.633124e+16 | 0 | 35 | 48 |